Software
Primer-Aided Truncation for the Creation of Hybrid Proteins (PATCHY)
Learn lessMany proteins consist of individual domains that are connected by so-called linker sequences with widely differing length, sequence and structure. In signal receptors, linkers connect the sensor unit that perceives cognate signals to the effector unit which elicits physiological responses. Signal transduction from sensor to effector hinges on the intervening linker; modifications in the linker as small as the exchange, the insertion or removal of a single amino acid residue can profoundly alter receptor output. For the design of signal receptors, this poses a particular challenge and often entails the construction and testing of multiple linker variants. To aid this process, we have developed the PATCHY method (Primer-Aided Truncation for the Creation of Hybrid Proteins) for the facile construction of defined gene libraries in which individual members differ in length and sequence of the linker between two gene fragments A and B.
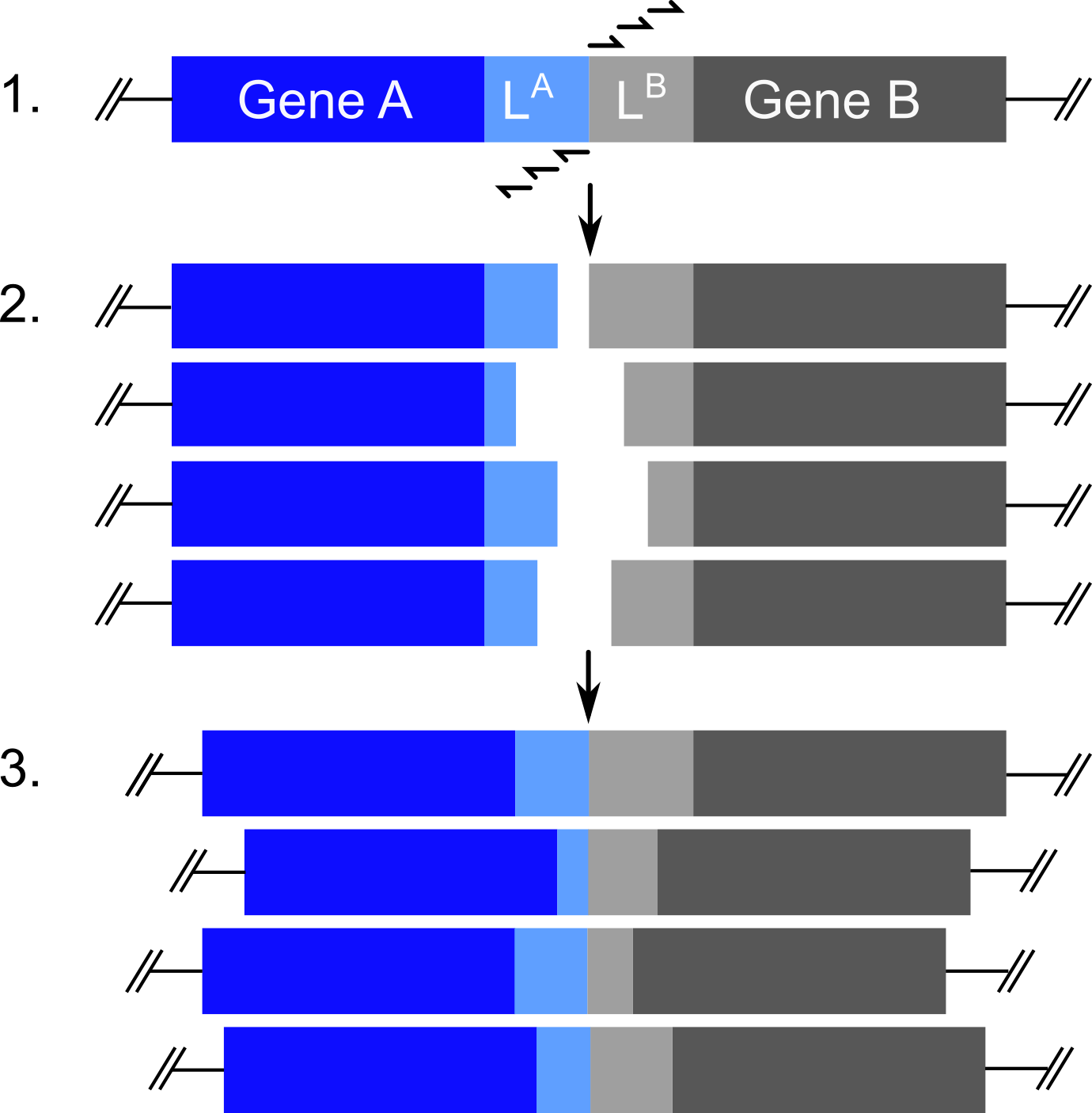 Figure - The PATCHY method generates gene libraries in which the gene fragments A and B
are connected by varying linkers of predefined length and composition. To this end, a precursor plasmid (step 1) in which fragments A and B are connected
by a linker of maximum length is amplified by PCR with sets of staggered forward and reverse primers. The resultant linear DNA fragments (step 2) are religated
to yield circular plasmids (step 3).
Figure - The PATCHY method generates gene libraries in which the gene fragments A and B
are connected by varying linkers of predefined length and composition. To this end, a precursor plasmid (step 1) in which fragments A and B are connected
by a linker of maximum length is amplified by PCR with sets of staggered forward and reverse primers. The resultant linear DNA fragments (step 2) are religated
to yield circular plasmids (step 3).
PATCHY is based on the amplification of a precursor plasmid with staggered primers as illustrated in the figure. We have developed a Python script which designs the required primers for PCR amplification. For details, cf. below references. To obtain the program, click on [Download].
- Ohlendorf, R., Schumacher, C.H., Richter, F., and Möglich, A. (2016). Library-Aided Probing of Linker Determinants in Hybrid Photoreceptors. ACS Synth. Biol. 5, 1117-1126. [Pubmed]
- Stabel, R., Stüven, B., Ohlendorf, R., and Möglich, A. (2017). Primer-Aided Truncation for the Creation of Hybrid Proteins. Meth. Mol. Biol. 1596, 287-304. [Pubmed]
Upgrading a Microplate Reader for Photobiology and All-optical Experiments
Learn lessAutomation can vastly reduce the cost of experimental labor and thus facilitate high experimental throughput, but little off-the-shelf hardware for the automation of illumination experiments is commercially available. Here, we use inexpensive open-source electronics to add programmable illumination capabilities to a multimode microplate reader. The setup is easily assembled and thus offers a facile approach to conducting illumination experiments at high throughput, reproducibility and fidelity.
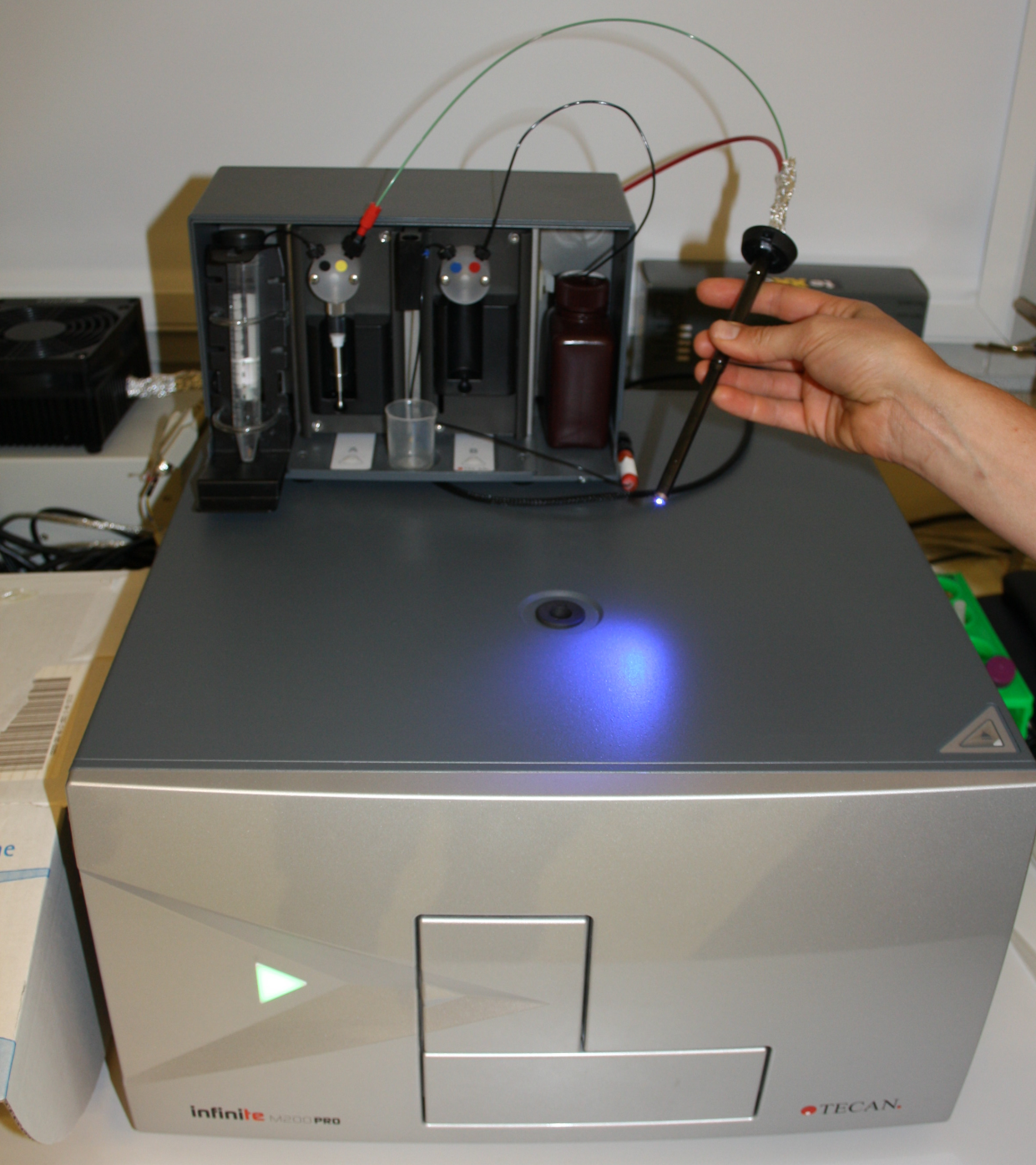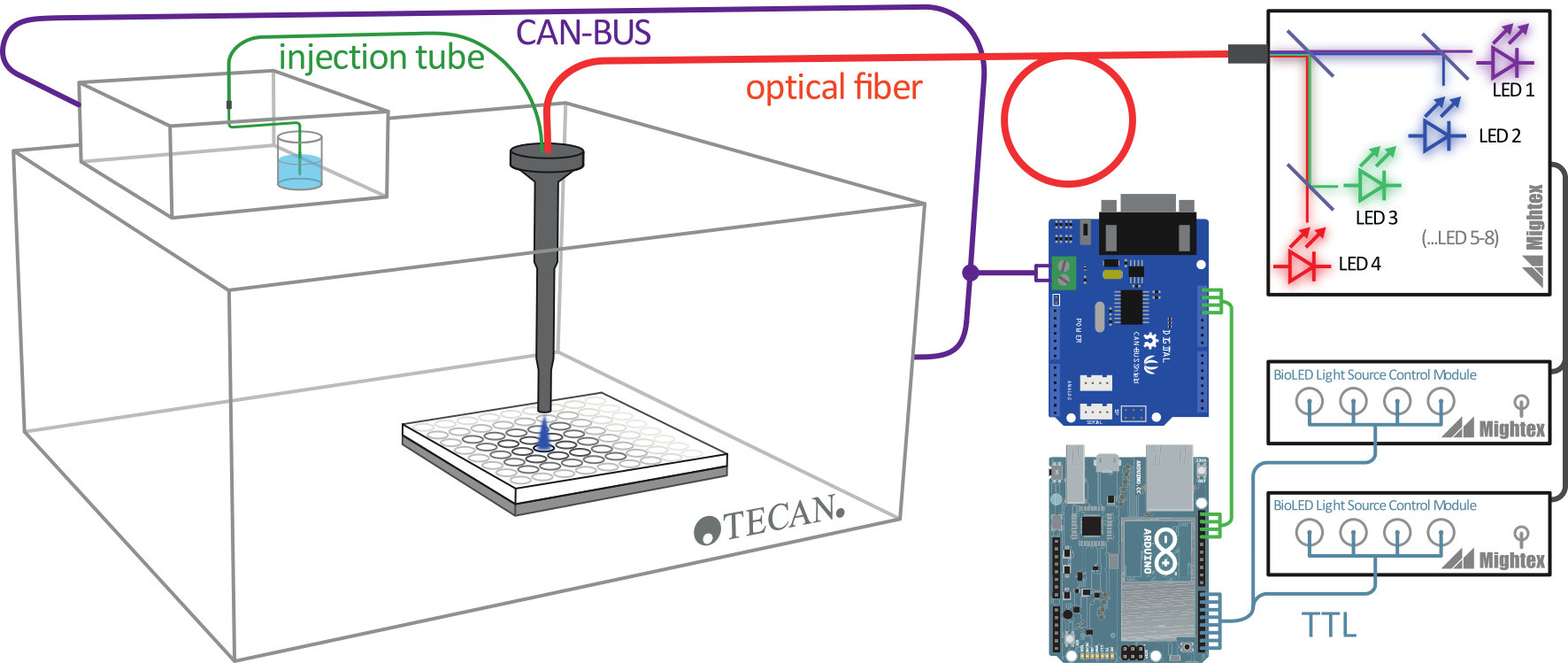
 Figure - To achieve programmable and online illumination of microtiter plates,
we intercepted the communication between a Tecan M200pro plate reader and its external injection device.
Using Arduino, we hijacked injection commands to control a LED light source. Light output from the LED was fed
into a fiber-optic guide and applied to the microtiter plate inside the instrument. (Disclaimer: Use at your own risk; know what you are doing.)
Figure - To achieve programmable and online illumination of microtiter plates,
we intercepted the communication between a Tecan M200pro plate reader and its external injection device.
Using Arduino, we hijacked injection commands to control a LED light source. Light output from the LED was fed
into a fiber-optic guide and applied to the microtiter plate inside the instrument. (Disclaimer: Use at your own risk; know what you are doing.)
For details, cf. below reference. To obtain the program, click on [Download].
- Richter, F., Scheib, U.S., Mehlhorn, J., Schubert, R., Wietek, J., Gernetzki, O., Hegemann, P., Mathes, T., and Möglich, A. (2015). Upgrading a microplate reader for photobiology and all-optical experiments. Photochem. Photobiol. Sci. 14, 270-279. [Pubmed]
Programmable LED Arrays for Photobiological Experiments
Learn lessExperiments in photobiology frequently demand the illumination of samples with light of specific wavelength and intensity for defined times. To facilitate such experiments, we have developed an Arduino-based, programmable LED array for the illumination of microtiter plates. Using a graphical user interface the color and intensity of an array of 8x8 LEDs can be configured.
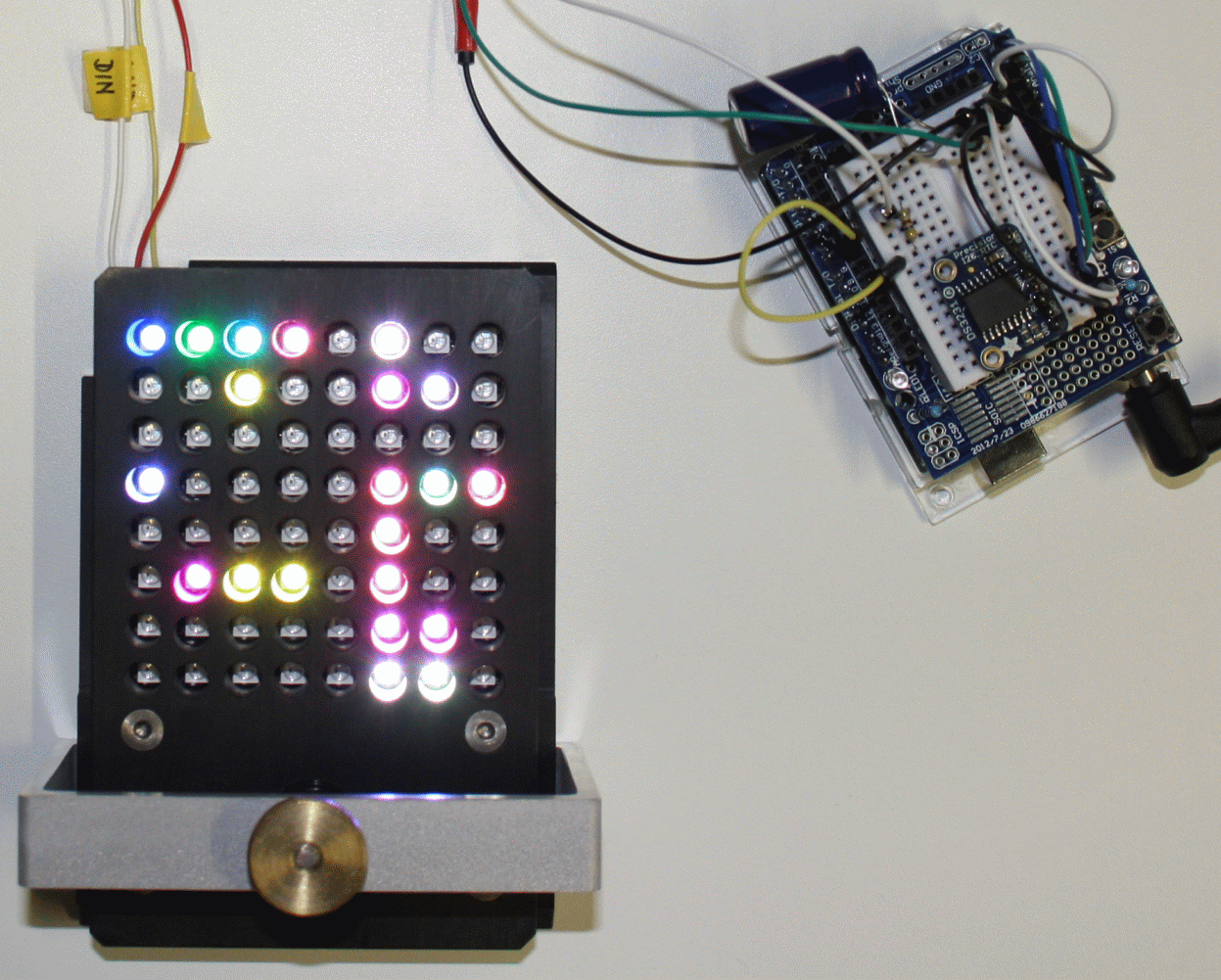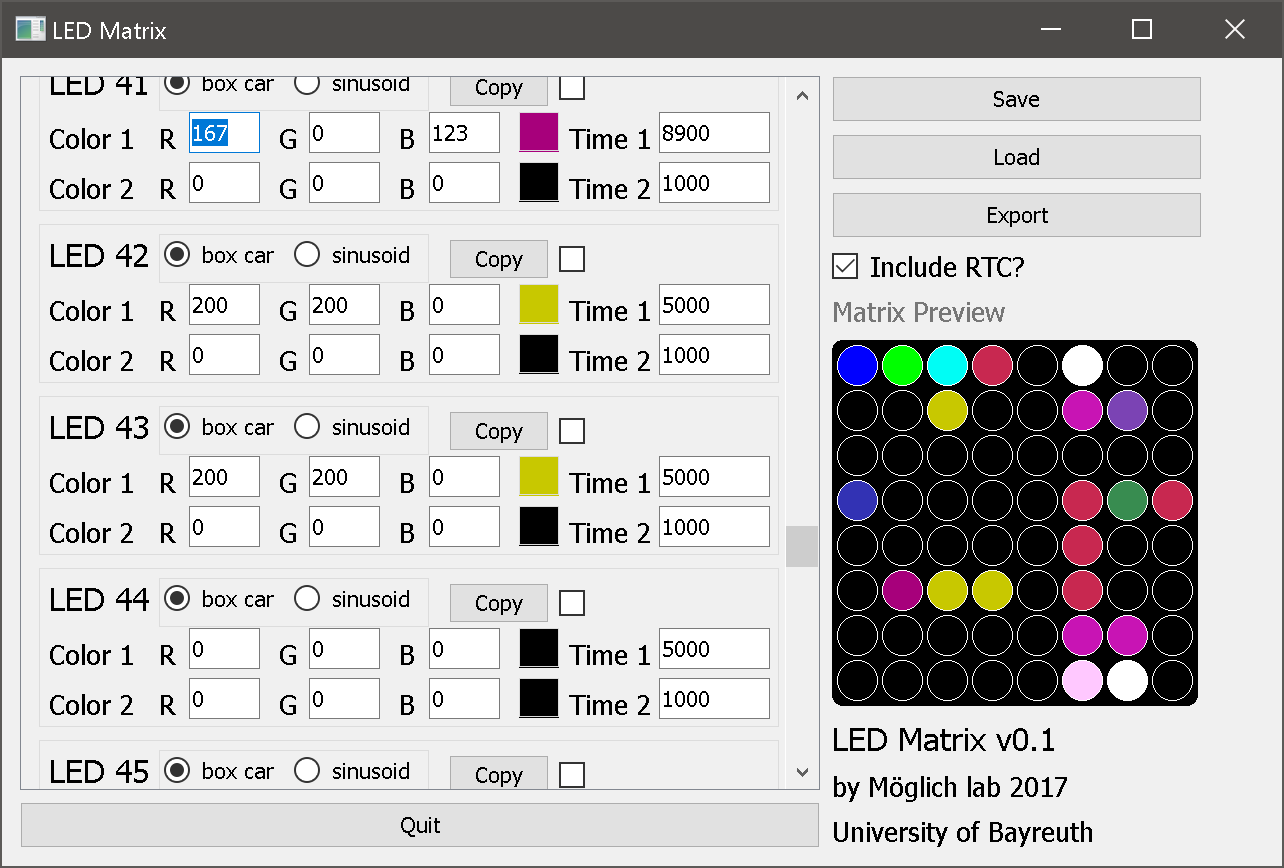
 Figure - A graphical user interface allows the easy configuration of an 8x8
LED matrix. The interface generates an Arduino sketch file for the control of the actual LED matrix.
Figure - A graphical user interface allows the easy configuration of an 8x8
LED matrix. The interface generates an Arduino sketch file for the control of the actual LED matrix.
Download program as [Python] or [Windows executable]. Download [version 1] or [version 2] of templates for 3D printing of adapter pieces. Download [Fritzing file] of Arduino setup.
The original LED array (above) used a commercially available LED matrix with emission in the blue, green and red regions of the visible spectrum. To enable experiments with photoreceptors responding to other light colors, we redesigned the setup to allow use of custom LEDs.
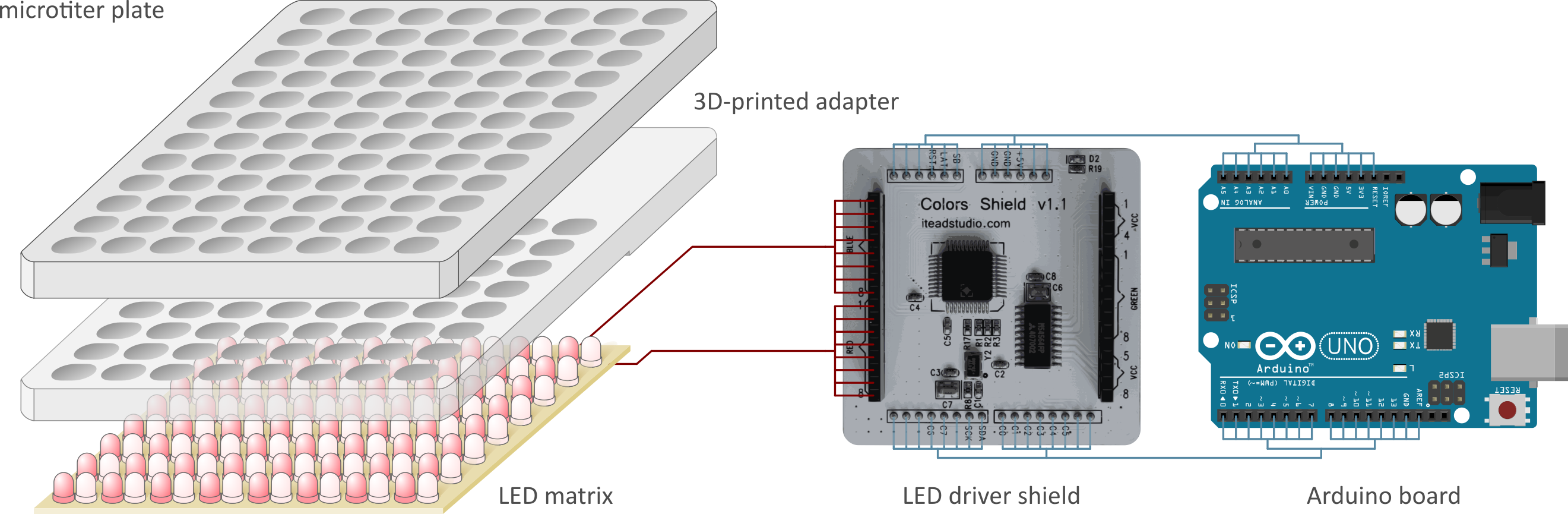 Figure - The reconfigured version of the programmable LED matrix allows the flexible use of custom LEDs, e.g., ones emitting in the red and far-red region of the electromagnetic spectrum.
Figure - The reconfigured version of the programmable LED matrix allows the flexible use of custom LEDs, e.g., ones emitting in the red and far-red region of the electromagnetic spectrum.
Download program graphical user interface as [Python] code. Download [3D-print templates] for matrix housing. Download [printed circuit board].
- Hennemann, J., Iwasaki, R.S., Grund, T.N., Diensthuber, R.P., Richter, F., and Möglich, A. (2018). Optogenetic Control by Pulsed Illumination. Chembiochem 19(12), 1296-1304. [Pubmed]
- Stüven, B., Stabel, R., Ohlendorf, R., Beck, J., Schubert, R., and Möglich, A. (2019). Characterization and engineering of photoactivated adenylyl cyclases. Biol Chem in press [doi]
Fit-o-mat, a Versatile Open-Source Program for Non-Linear Least-Squares Fitting
Learn lessFor detailed info, please visit the program page.
- Möglich, A. (2018). An open-source, cross-platform resource for nonlinear least-squares curve fitting. J Chem Educ 95(12), 2273-2278 [doi]

